The Institute Colloquium: Genomic scope of adaptive mutations in the face of an envir
Date
Monday, April 13, 2015 16:30 - 17:30
Speaker
Sally Otto (University of British Colombia)
Location
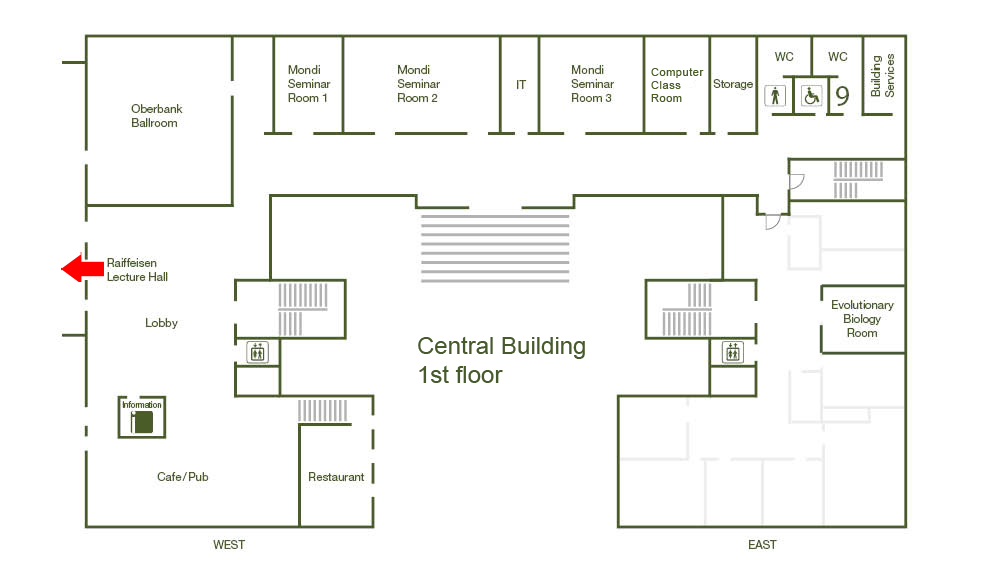
Raiffeisen Lecture Hall, Central Building
Series
Colloquium
Tags
Institute Colloquium
Contact
Url
As evolutionary biologists, we are only now getting a clear picture of the genetic basis of adaptation. Through genomic sequencing, we can pinpoint the mutations responsible for adaptation and use these mutations to parameterize models and to test evolutionary theory, providing a much sounder empirical basis for this theory. In this talk, I will describe experiments with Saccharomyces cerevisiae aimed at clarifying the nature and distribution of beneficial mutatioins. A panel of independent adaptive mutations was obtained by exposing haploid yeast to harsh environmental conditions: a fungicide (nystatin) or toxic levels of copper. The array of mutations allowing adaptation to the either environment was determined by genome-wide sequencing of 35 different adapted lines from each of the two environments. This panel was then used to measure trade-offs among environments and among ploidy levels, as well as properties such as dominance across environments. The implications for Haldane's sieve and the role of ploidy in adaptation will be discussed.