The Institute Colloquium: (Joint) segmentation and tracking with applications in biol
Date
Monday, May 4, 2015 16:30 - 17:30
Speaker
Fred Hamprecht (University of Heidelberg)
Location
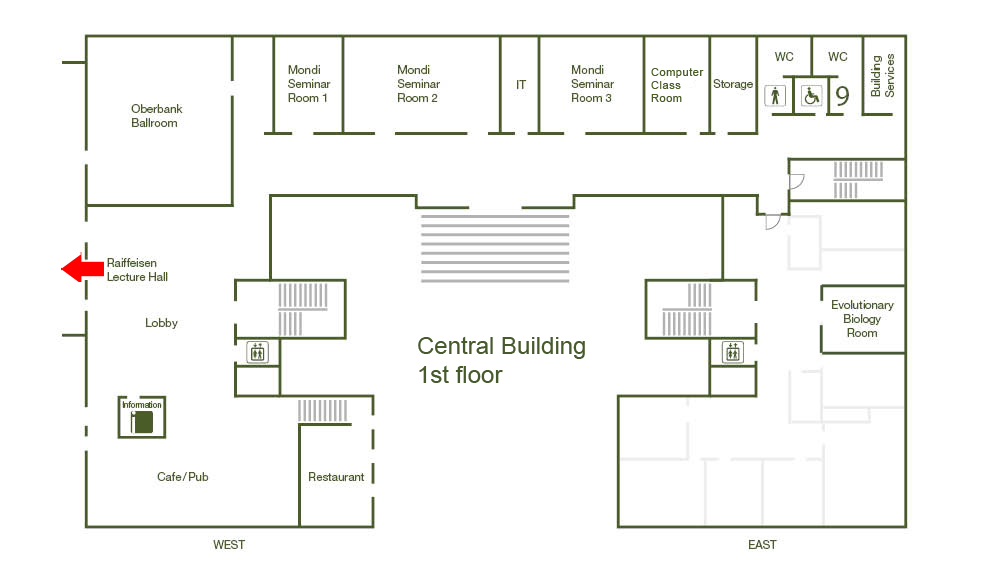
Raiffeisen Lecture Hall, Central Building
Series
Colloquium
Tags
Institute Colloquium
Contact
Url
The tracking of an unknown number of divisible cells, or of indistinguishable larvae, is an intriguing problem with important applications in developmental biology, ethology and beyond. I will show some of the best current approaches to solving this problem: firstly, a tracking-by-assignment approach that tries to detect all targets and then links them up while making allowance for undersegmentation, false positive detections and divisions. Secondly, I will present a model that addresses the detection and assignment steps jointly. Both formulations lead to difficult combinatorial optimization problems that can be solved to optimality for modestly sized instances. Finally, I will demo the former model, which is available as a workflow in the open source "ilastik" package; and will show a possible strategy to identify the less confident parts of a structured prediction for targeted proofreading.